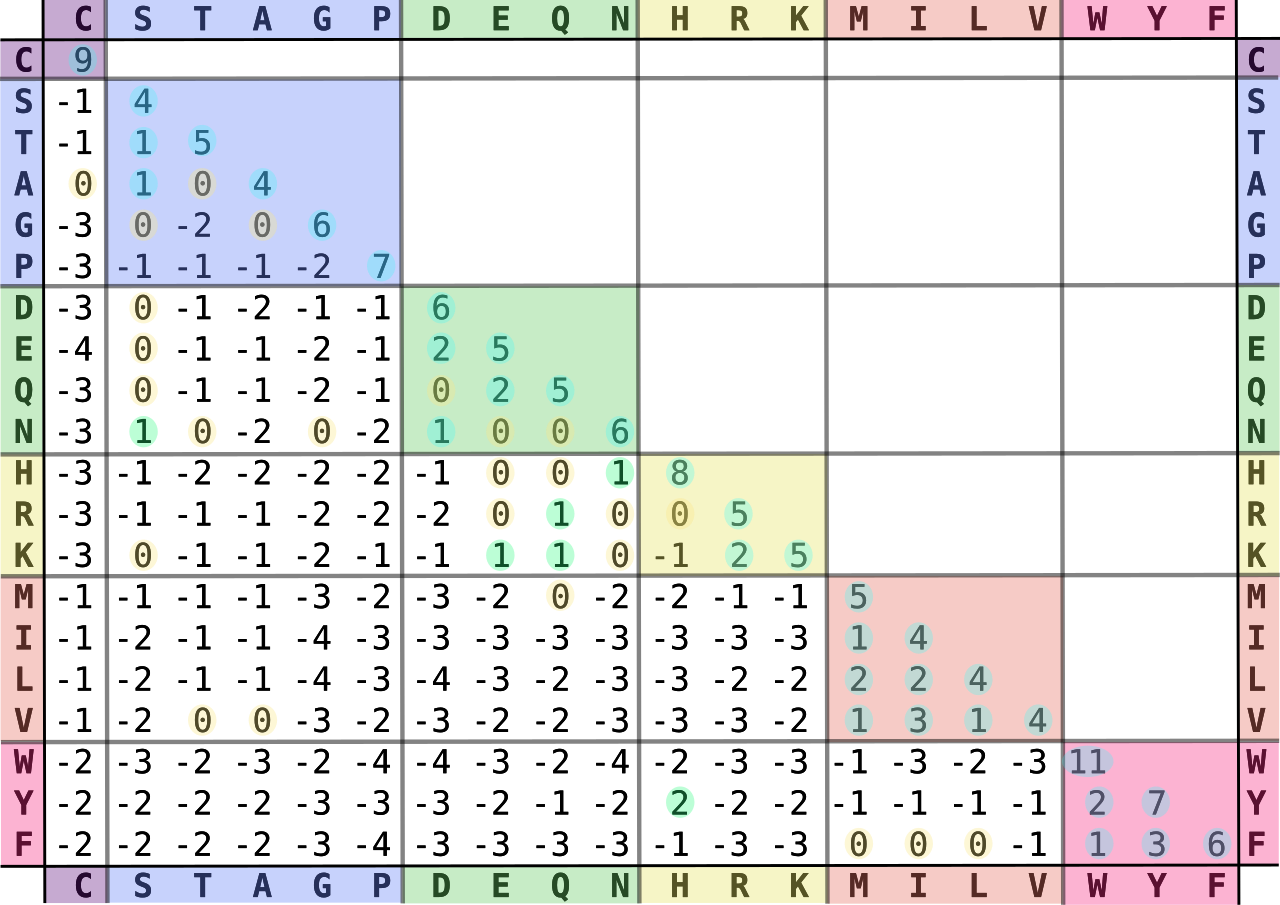
Multiple Sequence Alignment (MSA) is a process of aligning 2 or more protein, RNA or DNA sequences to maximize regions of sequence similarity. Gaps are used to align sequences with different lengths. Too many gaps can cause the alignment to become meaningless and so a gap penalty is used during the alignment process to ensure judicious usage of gaps. The final alignment score is computed by using substitution matrices. In the case of proteins this is done using BLOSUM (BLOcks SUbstitution Matrix) as shown in Figure 1. In practice, different substitution matrices are tailored to detecting similarities among sequences that are diverged by varying degrees. The BLOSUM-62 matrix, shown in Figure 1, is one of the best for detecting weak protein similarities.